Previously cross-posted at For Better Science
A recent paper by Li, Tomlenovic, Li & Shaw has attracted a great deal of attention, both positive and negative. The authors assert that aluminium compounds activate inflammation-related genes in the brains of mouse pups, causing brain damage, when they are injected with dosages comparable to the amounts contained in the US infant vaccination schedule.
In particular, the paper featured in a comment thread at Pubpeer, in the inimitable and inimical style that prevails in that forum. Here I will try to spin comments and observations from the thread into some sort of coherent narrative. I will not try to duplicate criticisms of the paper elsewhere, made by people more qualified than me, including Orac at Respectful Insolence and by Mad Virologist and The Blood-Brain Barrier Scientist... those critiques questioned whether the genes measured were an ideal selection, whether the semi-quantitative methods and statistics applied to them were appropriate, and whether the results of those methods (Figure 1, below) have been manipulated with Performance-Enhancing Photoshop. I will continue to spell "Aluminium" with two 'i's, the way God intended.
First, a little of the background is in order. The Journal of Inorganic Biochemistry (J.Inorg.Biochem) has a tradition of publishing papers on neurology, immunology, animal behaviour, and the causation of autism, as long as those topics arise in the context of aluminium toxicity. The Editors take a generous interpretation of their remit, and the journal's peer reviewers are optimistic about the bounds of their expertise. This tradition goes back at least to 2009. In 2011 the Editors surpassed themselves with a Special Issue ("Living the in Aluminum Age" [sic]), guest-edited by a high-ranking member of the Aluminati, Professor Christopher Exley of Keele University. Many surprising claims were made in that issue, such as the fact that alumina in antiperspirants is a leading cause of breast cancer, while Tomljenovic and Shaw contributed a problematic comparison between variations in autism prevalence and variations in vaccination schedules... a comparison that works best if one postulates that the neurotoxicity of aluminium adjuvants propagates backwards in time, in the manner of thiotimoline.
The paper of interest here, the most recent in this series, presents a comparison between RNA levels and protein levels in the brains of mice variously injected with colloidal aluminium hydroxide, or saline solution (Al and Control groups). The graphical component of the paper is dominated by RT-PCR gels (where short segments of RNA have been separated along a gel to mark their presence or absence after repeated cycles of reverse-transcriptase PCR), and Western blots. These are the white-on-black tandem kayak images and black-on-white moustache identity parades, respectively, in Figure 1.
Figure 1:
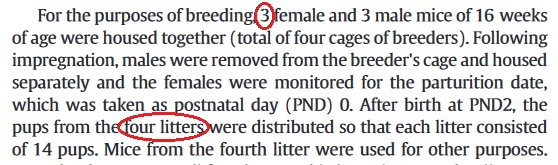
 Genetically-diverse CD-1 mice were used here -- "vulgar little mice" -- bred under conditions that are described in a manner that invites confusion, worded in terms of four groups rather than two. There is also uncertainty as to the number of breeding pairs.
Genetically-diverse CD-1 mice were used here -- "vulgar little mice" -- bred under conditions that are described in a manner that invites confusion, worded in terms of four groups rather than two. There is also uncertainty as to the number of breeding pairs.The description made a little more sense when it first appeared in a 2013 paper in J.Inorg.Biochem -- the first report from this specific project [https://www.ncbi.nlm.nih.gov/pubmed/23932735] -- in which we learn that there were two Al-exposed litters of 14 pups, with different exposure levels; also a fourth litter that was saved for feeding pet snakes or some other objective which need not concern us here. The ambivalence on the number of breeding mothers was present in 2013 as well.
The Al and Control mice received the same number of injections, on the same days... no, wait, according to the 2013 account, they received 6 and 5 injections respectively, on a different schedule.
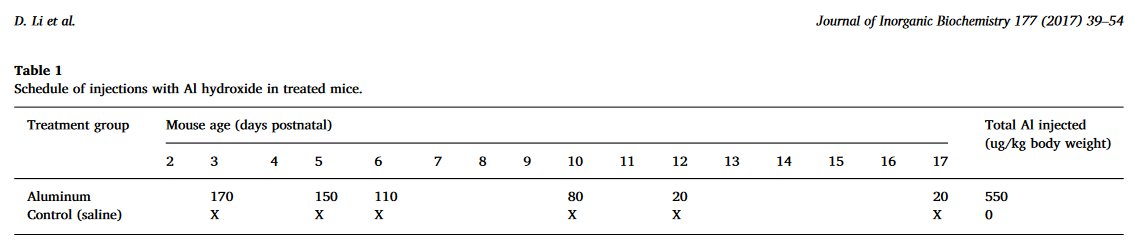
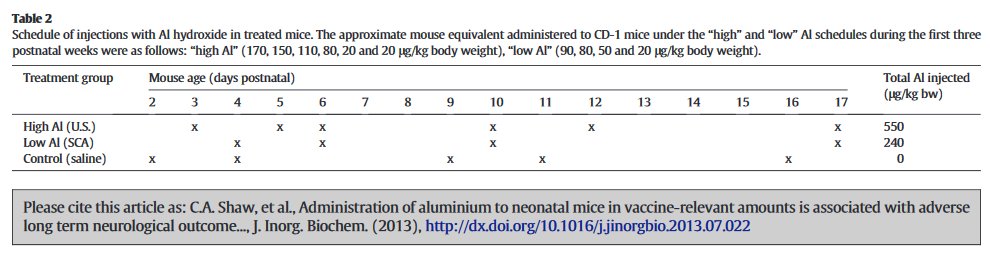
At the age of 16 weeks, having performed behavioural tasks for the edification of the experimenters, the mice were sacrificed on an altar of black basalt.
"At 16 weeks of age the mice were euthanized and the brain tissues were collected for gene expression profiling experiments."I made up the part about a basalt altar. Also, the mice survived to 34 weeks before their execution.
"The general development of mice was monitored by systematic recording of their weights from week 1 till the time of sacrifice (week 34)."I try to keep better track of the age at sacrifice of my experimental subjects, but it is true that I work with humans.
The posthumous adventures of those murine brains are best explained by an excursion outside the confines of J.Inorg.Biochem, to a pair of 2014 papers: "Are there negative CNS impacts of aluminum adjuvants used in vaccines and immunotherapy?" (Shaw, Li & Timljenovic), Immunotherapy, and "Etiology of autism spectrum disorders: Genes, environment, or both?" (Shaw, Sheth, Li & Timljenovic), OA Autism.* Both contain foretastes of Figure 1, as a kind of appetite-whetting trailer for the present paper **:
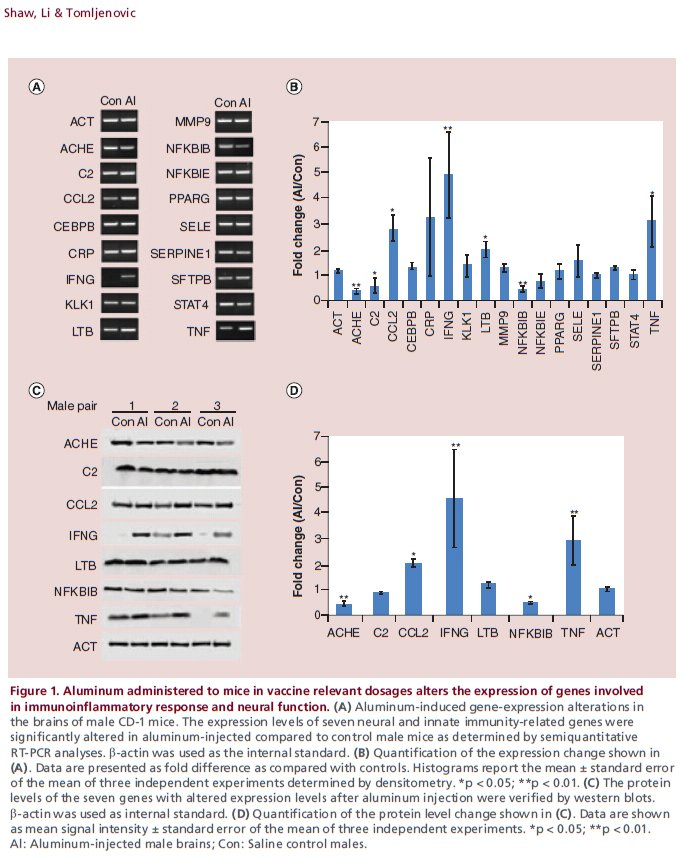

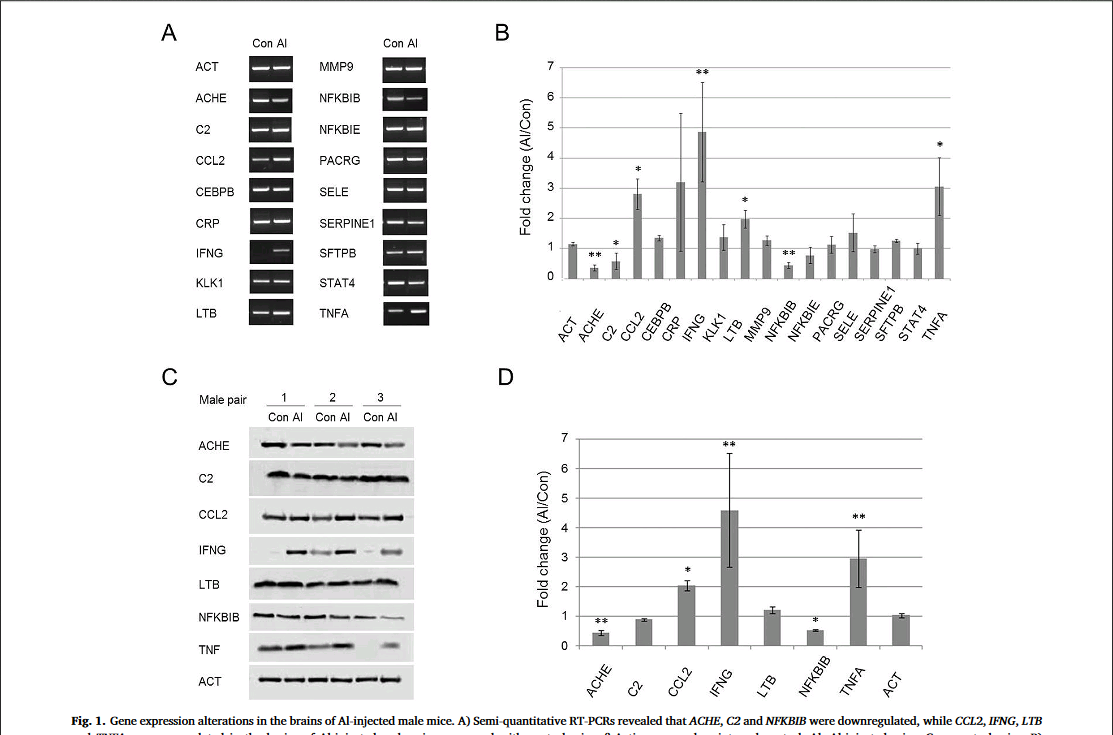
Six male mice were chosen for preliminary measurements: three Al-exposed and three controls. It is not stated whether they were selected at random, or because their behavioural performance promised results that would fit the desired picture. Both 2014 papers included a caveat that the samples were too small to sustain strong conclusions, and the results were merely suggestive until more comprehensive measurements can be made. In fact the reader might wonder how the confidence bounds shown in the various Figures 1 can be drawn at all from only three measurements.


So the 2017 paper is the fulfilment of that promise to "collect the data on other samples". There are now 10 male mice involved, and 10 females:
The brain samples of five males and five females injected with Al and five males and five female control mice were randomly paired for gene expression profilingIn addition,
The experiments for each mouse were repeated three times for statistical purposes... Perhaps under the impression that these would be independent measurements, and could be used to narrow the confidence limits.
But mirabile dictu, the 2017 Figure 1 (see above) is exactly the same as the 2014 versions! -- neither the mean values nor the error bars for male-mouse brains were affected by the additional data! This brought consternation and speculation into the Pubpeer thread.
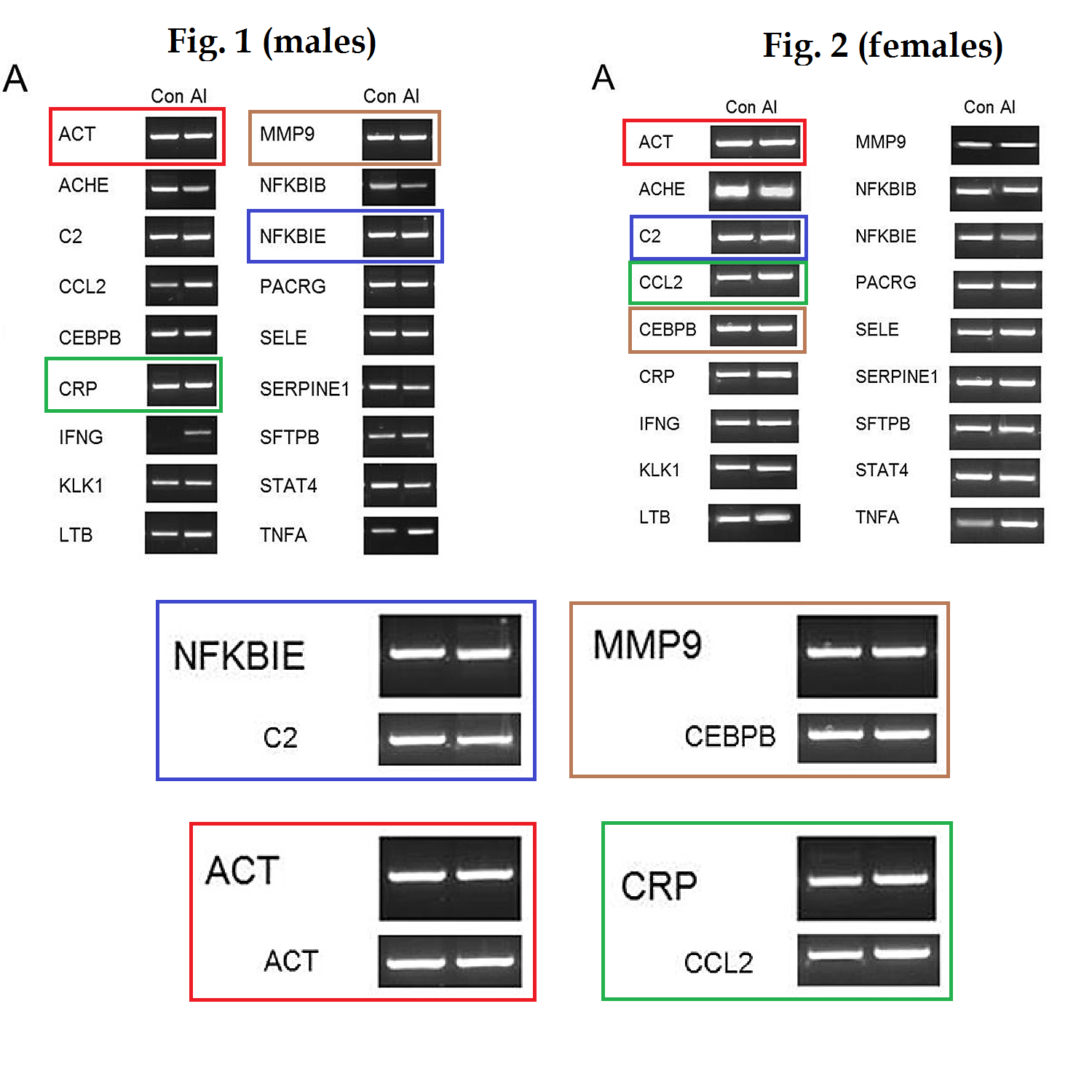
In the course of that long history of previous publication, the panels comprising Figure 1 have undergone multiple cycles of contrast enhancement and high-loss JPEG image compression, so one can download a "high resolution" version but the main benefit is to see the compression artefacts at greater magnification. The present paper also added corresponding figures for the female-mouse brains, notably Figure 2.
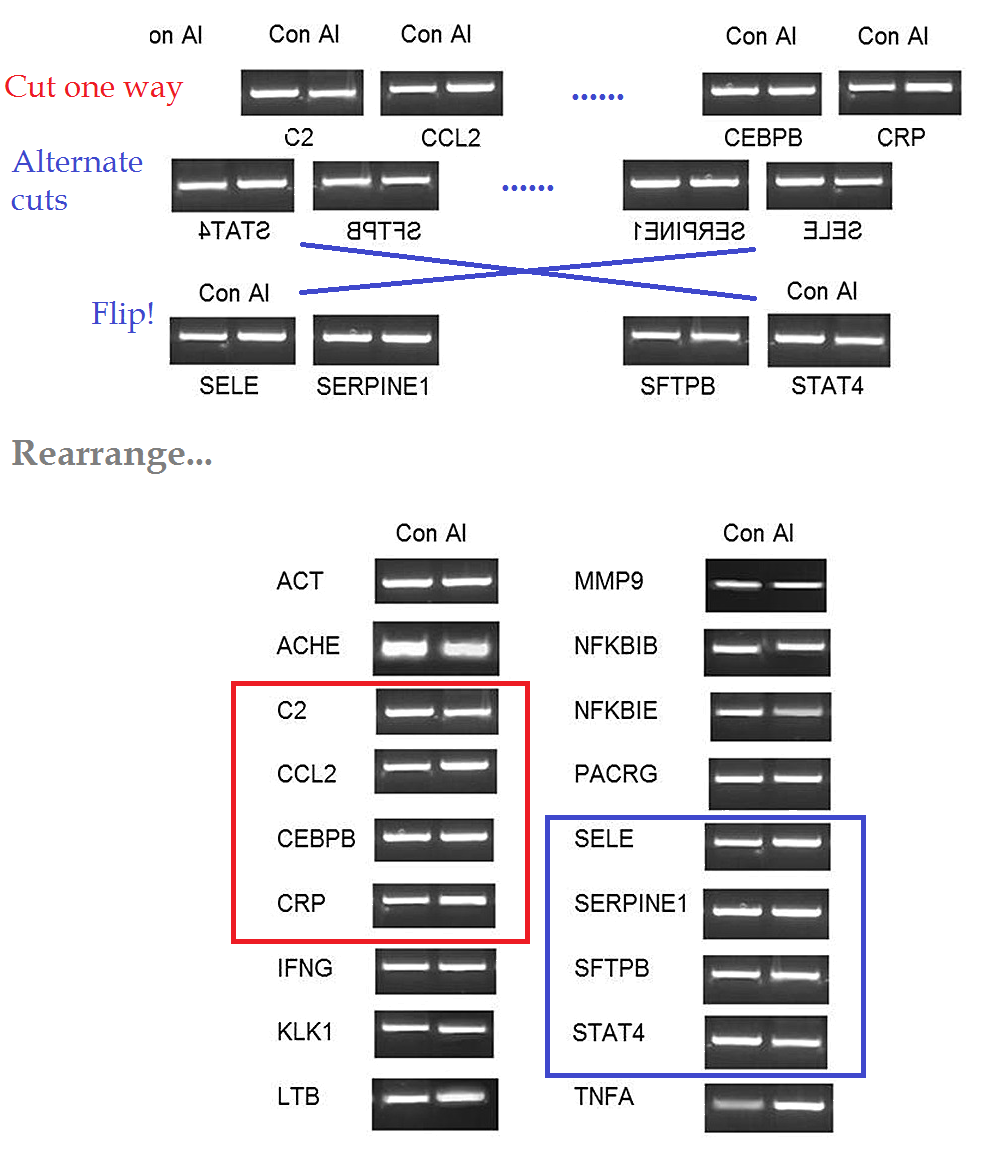
Further consternation resulted when sharp-eyed Pubpeer contributors noted that some of the RT-PCR kayaks are distinctive in appearance -- one boasting a little foresail (the Control C2 RNA) while another carries a little-man-in-the-boat as its passenger (Al CEBPB RNA) -- and these allow one to recognise that several of the Al-Control pairs had previously been extracted from the brains of male mice (where they coded for the NFKBIE and MMP9 proteins respectively).
Nor does it stop there. Several of these distinctive kayaks have been reflected left-to-right -- so the foresailed band that evinced the presence of C2 RNA (in the left-hand column of Figure 2A) appears again (in the right-hand column), heading in the other direction, to evince the presence of SFTPB. The little-man-in-the-boat reappears in the same way as SERPINE1, while a kayak with an outboard motor -- previously a Control for CRP -- becomes the Control for SELE.
It is as if someone had been told to turn data sets Y and Z into figures for the paper, but those data did not exist, so the team member improvised from data set X, already published. Or else they thought they were studying mirror neurons. In total the same images have appeared in five places, signifying three different protein RNAs, which shows admirable economy on someone's part.
That is still not all; the horizontal-mirror methodology was applied again to Figure 4.
Certain bands in these RT-PCR gels are marked by distinctive bubbles and hiccoughs. These allow us to recognise that Figure 4D is a reflected version of 4B, differently exposed. Kayaks displaying the presence of Cytokine Actin RNA in 4B became evidence for Adhesion-molecule ICAM-1 in 4D (while IL-6 became E-Selectin, IL-5 became VCAM-1 and IL-4 became Actin).
I am no expert on the minutiae of PCR methods but I am fairly sure that inverting images -- reversing the status of Control and Al-exposed genetic activity -- is not legitimate, and it does not happen by accident.
Footnotes
* OA Autism comes from a one-man publisher "OA London Publishing", a source of interest and entertainment in its own right, as reported a few years ago by Neuroskeptic. The journal is moribund, with no new additions to its files since late 2014, but the website is still there so we should enjoy it while it lasts. Evidently the publisher managed to recruit Alexander Seifalian to be Chair of the "Board of Trustees" as a token of credibility: his name may be familiar to Leonid's readers.
** Hat tip to Michael Brown for spotting the foreshadowing.





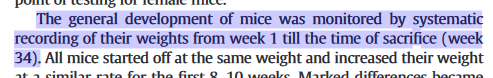




2 comments:
"The experiments for each mouse were repeated three times for statistical purposes"
How does one euthanatize a mouse three times? One would think they'd get chary. Or perhaps they were cats.
I think the "repeated three times for each mouse" wibble was an act of creative misreading, or poetic Deconstructivist misprision, on the part of the authors. In 2013 the results had been described as coming from three mice (or rather, three control / treated pairs of mice). Now the authors have retconned the same values and figures as representing five pairs of mice. But the original claim about "three observations" crept back into the text, as a dim recollection or as some sort of textual inertia, now manifesting as a completely spurious report of three repetitions per mouse.
It is hard to be sure because none of the authors will admit to writing the words of the paper.
Post a Comment